Teaching, side quests, writings, and outreaches
Teaching
Youtube tutorials
When there are enough storage space in my little laptop, I create tutorial videos on youtube.
An Invitation for Horticultural Students to Bioinformatics
I delivered a mini course about transcriptome analysis as a guest lecturer in the course “Methodology in Horticulture Crop (IV)” at NTU (National Taiwan University). By launching a Facebook campaign page, I got twice participants than expected.
Prerequisite Learning List and Resources
- Next-generation sequencing
- Introduction to molecular biology
- Process of transcription
- Preparing your development environment
- Annotation systems
- Biostatistics
Slides and Handout
Video Slideshow is available. Raw Powerpoint file can be obtained through DM or from chatbot of RNA-Sick Facebook page:
https://m.me/rna.sick?ref=Welcome
Handout
Writings
Quick Guide to Bioinformatics for biology background students (in Chinese)
In this quick guide, I intended to bypass the technical details, try to focus on the building of self-teaching habits, the value of academic integrity and growth mindset. Once the readers get an overview of what should be expected to learn, they should go on and acquire new knowledge from primary sources such as original research articles published by international academic journals. You can view the original version here on IT Help. It is consists of four modules:
Introduction and minimal computer skill sets
Assuming that readers have a biology/horticulture background, some computer utilization skills are required to start the journey into bioinfomatics. Just to list a few: familirity with unix-like environment and using command-line, text editor setting, running scripts (maybe R or Python).
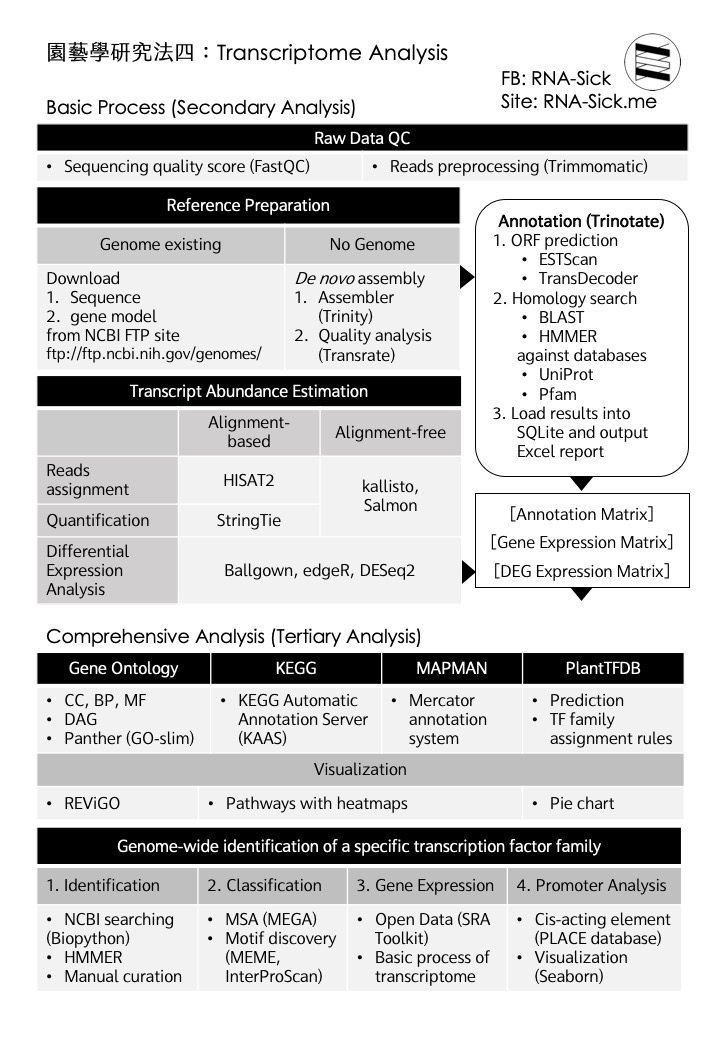
Transcriptome pipeline
RNA-Sequencing (RNA-Seq) is one of the major game changing technologies in crop research. It is also the main reason for normal biology/horticulture students to further dive into bioinformatics. Although most of the sequencing service providers nowadays would generate a basic report containing figures and tables that can be directly used for publication, understanding the principles and algorithms is still a must for correct interpretation.
Comprehensive Analysis
After acquiring differentially expressed genes from RNA-Seq, there are various ways to discover biological meanings from the data. Interesting results can be yielded by using heatmap to visualize transcript abundance, gene clustering according to expression pattern, or accessing online database using custom script.
Road to an Academic Career
Besides the technical materials regarding data processing or sequence analysis, I included a part to talk about how to make use of digital tools to strengthen academic integrity and build personal brand as early-career researcher.
Paper Digest
I translate, summarize papers and post them here (mostly in Chinese).

Social Media
I am posting Bioinformatic tips and memes on these social media:
Facebook: https://www.facebook.com/rna.sick Instagram: https://www.instagram.com/rna_sick/ Twitter: https://twitter.com/rna_sick